Time Series Forecasting Lab (Part 6) - Stacked Ensembles

Cover photo by Markus Spiske on Unsplash
Go to R-bloggers for R news and tutorials contributed by hundreds of R bloggers.
Introduction
This is the sixth of a series of 6 articles about time series forecasting with panel data and ensemble stacking with R.
Through these articles I will be putting into practice what I have learned from the Business Science University training course 1 DS4B 203-R: High-Performance Time Series Forecasting", delivered by Matt Dancho. If you are looking to gain a high level of expertise in time series with R I strongly recommend this course.
The objective of this article is to learn how to build multi-level stacking ensembles with modeltime. In the figure below we start from the bottom by reminding us of each individual model or submodel performance from Part 4 (non-tuned models) and Part 5 (tuned models). Then we will define meta learners which learn from k-fold cross-validation predictions from all submodels. We can compare meta-learners' performance and select best meta-learners to be used for defining a weighted average ensemble to produce a final multi-level stacking model.

Prerequisites
I assume you are already familiar with the following topics, packages and terms:
dplyr or tidyverse R packages
k-fold cross-validation
hyperparameter tuning from Part 4
average and weighted ensembles from Part 5
Packages
The following packages must be loaded:
# 01 FEATURE ENGINEERING
library(tidyverse) # loading dplyr, tibble, ggplot2, .. dependencies
library(timetk) # using timetk plotting, diagnostics and augment operations
library(tsibble) # for month to Date conversion
library(tsibbledata) # for aus_retail dataset
library(fastDummies) # for dummyfying categorical variables
library(skimr) # for quick statistics
# 02 FEATURE ENGINEERING WITH RECIPES
library(tidymodels) # with workflow dependency
# 03 MACHINE LEARNING
library(modeltime) # ML models specifications and engines
library(tictoc) # measure training elapsed time
# 04 HYPERPARAMETER TUNING
library(future)
library(doFuture)
library(plotly)
# 05 ENSEMBLES
library(modeltime.ensemble)
Reminder of individual models
Leu us remind us of the performance of each individual model or submodel. We do not consider the ensemble models from Part 5.
# Load all calibration tables (tuned & non-tuned models)
> calibration_tbl <- read_rds("workflows_NonandTuned_artifacts_list.rds")
> calibration_tbl <- calibration_tbl$calibration
> calibration_tbl %>%
modeltime_accuracy() %>%
arrange(rmse)
# A tibble: 8 x 9
.model_id .model_desc .type mae mape mase smape rmse rsq
<int> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 7 PROPHET W/ REGRESSORS Test 0.189 21.6 0.417 21.5 0.240 0.856
2 3 PROPHET W/ REGRESSORS - Tuned Test 0.189 21.6 0.418 21.5 0.241 0.857
3 4 PROPHET W/ XGBOOST ERRORS - Tuned Test 0.191 25.7 0.421 19.9 0.253 0.780
4 8 PROPHET W/ XGBOOST ERRORS Test 0.204 24.8 0.451 20.8 0.287 0.753
5 2 XGBOOST - Tuned Test 0.219 23.6 0.482 22.9 0.296 0.762
6 6 XGBOOST Test 0.216 25.0 0.476 22.4 0.299 0.747
7 1 RANGER - Tuned Test 0.231 26.1 0.509 24.6 0.303 0.766
8 5 RANGER Test 0.235 25.4 0.519 25.0 0.312 0.765
Stacking Algorithms
A stacking algorithm or meta-learner learns from predictions. The predictions come from k-fold cross-validations applied to each submodel. Once fed into the meta-learner tunable specification a hyperpameter tuning will be peformed for the meta-learner.
Generate predictions for meta-learners
Remember, for non-sequential models you must perform a k-fold cross-validation and not a time series cross-validation. Let us get resampeld predictions with modeltime_fit_resamples(). Here, k=10 folds.
set.seed(123)
resamples_kfold <- training(splits) %>%
drop_na() %>%
vfold_cv(v = 10)
tic()
submodels_resamples_kfold_tbl <- calibration_tbl %>%
modeltime_fit_resamples(
resamples = resamples_kfold,
control = control_resamples(
verbose = TRUE,
allow_par = TRUE,
)
)
toc()
Setup parallel processing
Parallel processing is set and toggled on by running the following commands:
# Parallel Processing ----
registerDoFuture()
n_cores <- parallel::detectCores()
plan(
strategy = cluster,
workers = parallel::makeCluster(n_cores)
)
Meta-learners specification
In this article we will use 3 meta-learners: Random Forest, XGBoost, and SVM.
Create a stacking algorithm specification with ensemble_model_spec(). Within the meta-learner specification we must provide the tunable parameters, the number of folds, and the size of the grid. We will also set grid search control parameters by activating verbose and allowing parallel processing.
For all meta-learners specifications we will define a k=10 folds and a grid size of 20.
Random Forest stacking algorithm
We perform tuning on 2 parameters: mtry and min_n.
We notice that the in-sample RMSE for the meta-learner is 0.125. Its performance on test data has RMSE = 0.230 and RSQ = 0.817.
tic()
set.seed(123)
ensemble_fit_ranger_kfold <- submodels_resamples_kfold_tbl %>%
ensemble_model_spec(
model_spec = rand_forest(
trees = tune(),
min_n = tune()
) %>%
set_engine("ranger"),
kfolds = 10,
grid = 20,
control = control_grid(verbose = TRUE,
allow_par = TRUE)
)
toc()
i Model Parameters:
# A tibble: 1 x 8
trees min_n .metric .estimator mean n std_err .config
<int> <int> <chr> <chr> <dbl> <int> <dbl> <chr>
1 1906 32 rmse standard 0.176 10 0.00225 Preprocessor1_Model02
i Prediction Error Comparison:
# A tibble: 9 x 3
.model_id rmse .model_desc
<chr> <dbl> <chr>
1 1 0.196 RANGER - Tuned
2 2 0.183 XGBOOST - Tuned
3 3 0.207 PROPHET W/ REGRESSORS - Tuned
4 4 0.179 PROPHET W/ XGBOOST ERRORS - Tuned
5 5 0.203 RANGER
6 6 0.203 XGBOOST
7 7 0.208 PROPHET W/ REGRESSORS
8 8 0.207 PROPHET W/ XGBOOST ERRORS
9 ensemble 0.125 ENSEMBLE (MODEL SPEC)
modeltime_table(
ensemble_fit_xgboost_kfold
) %>%
modeltime_accuracy(testing(splits))
# A tibble: 1 x 9
.model_id .model_desc .type mae mape mase smape rmse rsq
<int> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 1 ENSEMBLE (RANGER STACK): 8 MODELS Test 0.166 21.8 0.367 18.2 0.230 0.817
XGBoost stacking algorithm
We perform tuning on 4 parameters: trees, tree_depth, learn_rate, loss_reduction.
We notice that the in-sample RMSE for the meta-learner is 0.0957. Its performance on test data RMSE = 0.251 and RSQ = 0.769.
tic()
set.seed(123)
ensemble_fit_xgboost_kfold <- submodels_resamples_kfold_tbl %>%
ensemble_model_spec(
model_spec = boost_tree(
trees = tune(),
tree_depth = tune(),
learn_rate = tune(),
loss_reduction = tune()
) %>%
set_engine("xgboost"),
kfolds = 10,
grid = 20,
control = control_grid(verbose = TRUE,
allow_par = TRUE)
)
toc()
# A tibble: 1 x 10
trees tree_depth learn_rate loss_reduction .metric .estimator mean n std_err .config
<int> <int> <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
1 1717 8 0.0330 0.00876 rmse standard 0.180 10 0.00175 Preprocessor1_Model10
i Prediction Error Comparison:
# A tibble: 9 x 3
.model_id rmse .model_desc
<chr> <dbl> <chr>
1 1 0.196 RANGER - Tuned
2 2 0.183 XGBOOST - Tuned
3 3 0.207 PROPHET W/ REGRESSORS - Tuned
4 4 0.179 PROPHET W/ XGBOOST ERRORS - Tuned
5 5 0.203 RANGER
6 6 0.203 XGBOOST
7 7 0.208 PROPHET W/ REGRESSORS
8 8 0.207 PROPHET W/ XGBOOST ERRORS
9 ensemble 0.0957 ENSEMBLE (MODEL SPEC)
> modeltime_table(
ensemble_fit_xgboost_kfold
) %>%
modeltime_accuracy(testing(splits))
# A tibble: 1 x 9
.model_id .model_desc .type mae mape mase smape rmse rsq
<int> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 1 ENSEMBLE (XGBOOST STACK): 8 MODELS Test 0.182 23.9 0.401 19.7 0.251 0.769
SVM stacking algorithm
We perform tuning on 3 parameters: cost, rbf_sigma, margin. Run ?svm_rbf for explanations.
We notice that the in-sample RMSE for the meta-learner is 0.172. Its performance on test data RMSE = 0.219 and RSQ = 0.823.
tic()
set.seed(123)
ensemble_fit_svm_kfold <- submodels_resamples_kfold_tbl %>%
ensemble_model_spec(
model_spec = svm_rbf(
mode = "regression",
cost = tune(),
rbf_sigma = tune(),
margin = tune()
) %>%
set_engine("kernlab"),
kfold = 10,
grid = 20,
control = control_grid(verbose = TRUE,
allow_par = TRUE)
)
toc()
# A tibble: 1 x 9
cost rbf_sigma margin .metric .estimator mean n std_err .config
<dbl> <dbl> <dbl> <chr> <chr> <dbl> <int> <dbl> <chr>
1 4.60 0.0110 0.0595 rmse standard 0.172 10 0.00222 Preprocessor1_Model07
i Prediction Error Comparison:
# A tibble: 9 x 3
.model_id rmse .model_desc
<chr> <dbl> <chr>
1 1 0.196 RANGER - Tuned
2 2 0.183 XGBOOST - Tuned
3 3 0.207 PROPHET W/ REGRESSORS - Tuned
4 4 0.179 PROPHET W/ XGBOOST ERRORS - Tuned
5 5 0.203 RANGER
6 6 0.203 XGBOOST
7 7 0.208 PROPHET W/ REGRESSORS
8 8 0.207 PROPHET W/ XGBOOST ERRORS
9 ensemble 0.172 ENSEMBLE (MODEL SPEC)
> modeltime_table(
ensemble_fit_svm_kfold
) %>%
modeltime_accuracy(testing(splits))
# A tibble: 1 x 9
.model_id .model_desc .type mae mape mase smape rmse rsq
<int> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 1 ENSEMBLE (KERNLAB STACK): 8 MODELS Test 0.160 21.8 0.352 18.0 0.219 0.823
Multi-Level Stacks
The previous stacking approaches can be mutualized in a multi-level stack. We can combine meta-learners into a higher level of the stack which will be a normal (weighted) average ensemble as introduced in Part 5.
Below is a reminder of the previous stacking algorithms' performance:
modeltime_table(
ensemble_fit_ranger_kfold,
ensemble_fit_xgboost_kfold,
ensemble_fit_svm_kfold
) %>%
modeltime_accuracy(testing(splits)) %>%
arrange(rmse)
# A tibble: 3 x 9
.model_id .model_desc .type mae mape mase smape rmse rsq
<int> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 3 ENSEMBLE (KERNLAB STACK): 8 MODELS Test 0.160 21.8 0.352 18.0 0.219 0.823
2 1 ENSEMBLE (RANGER STACK): 8 MODELS Test 0.166 21.8 0.367 18.2 0.230 0.817
3 2 ENSEMBLE (XGBOOST STACK): 8 MODELS Test 0.182 23.9 0.401 19.7 0.251 0.769
Let us now define a weighted average ensemble model from the stacking algorithms as we did in Part 5. We will apply the ranking technique to calculate the weights.
First we create the accuracy results table on which we apply the ranking technique for weigths calculation, then we create the weighted average ensemble model with ensemble_weighted() by applying the weights, and finally, we evaluate the model.
loadings_tbl <- modeltime_table(
ensemble_fit_ranger_kfold,
ensemble_fit_xgboost_kfold,
ensemble_fit_svm_kfold
) %>%
modeltime_calibrate(testing(splits)) %>%
modeltime_accuracy() %>%
mutate(rank = min_rank(-rmse)) %>%
select(.model_id, rank)
stacking_fit_wt <- modeltime_table(
ensemble_fit_ranger_kfold,
ensemble_fit_xgboost_kfold,
ensemble_fit_svm_kfold
) %>%
ensemble_weighted(loadings = loadings_tbl$rank)
> stacking_fit_wt %>%
modeltime_calibrate(testing(splits)) %>%
modeltime_accuracy() %>%
arrange(rmse)
# A tibble: 1 x 9
.model_id .model_desc .type mae mape mase smape rmse rsq
<int> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 1 ENSEMBLE (WEIGHTED): 3 MODELS Test 0.161 21.8 0.354 18.1 0.223 0.817
With RMSE = 0.223, the stacked ensemble algorithm outperformed all algorithms!
Forescast plot
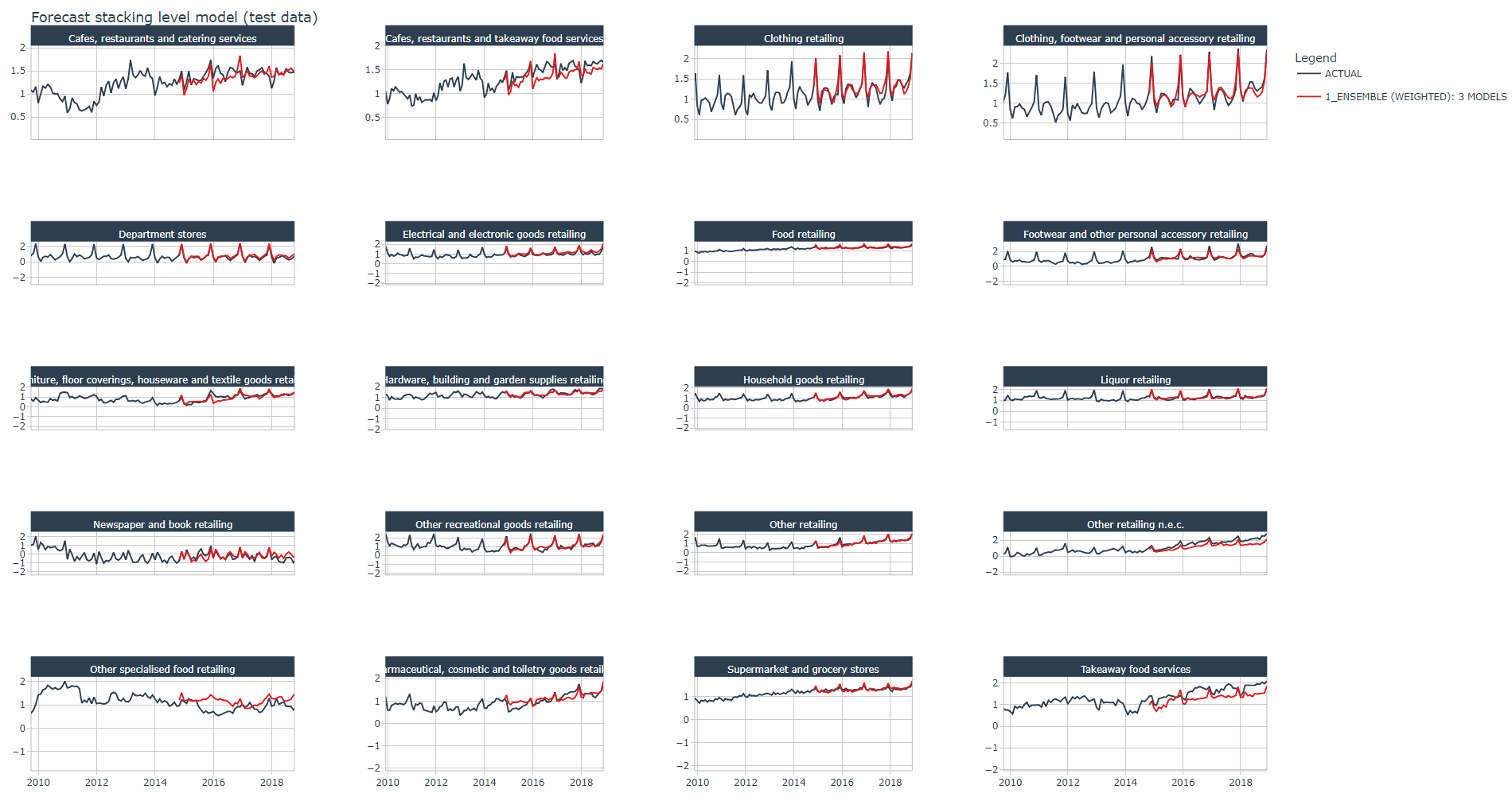
Below the stacked ensemble algorithm's forecast plot on the test dataset for all industries, it looks good ! except for one industry the model seems to follow pretty well the trend and spikes of the actual data.
calibration_stacking <- stacking_fit_wt %>%
modeltime_table() %>%
modeltime_calibrate(testing(splits))
calibration_stacking %>%
modeltime_forecast(
new_data = testing(splits),
actual_data = artifacts$data$data_prepared_tbl,
keep_data = TRUE
) %>%
group_by(Industry) %>%
plot_modeltime_forecast(
.facet_ncol = 4,
.conf_interval_show = FALSE,
.interactive = TRUE,
.title = "Forecast stacking level model (test data)"
)
Next 12 months Turnover forecast
Let us now (finally!) perform the next 12 months Turnover forecast for all 20 industries as we promised in Part 1.
First, we must refit the stack-level weighted average model with the prepared dataset from Part 2. We do this with modeltime_refit().
Then we calculate the Turnover predictions over the next 12 months (values are still transformed). We do this with modeltime_forecast() and by setting new_data to the future dataset.
# Toggle ON parallel processing
plan(
strategy = cluster,
workers = parallel::makeCluster(n_cores)
)
# Refit the model on prepared dataset
tic()
set.seed(123)
refit_stacking_tbl <- calibration_stacking %>%
modeltime_refit(
data = artifacts$data$data_prepared_tbl,
resamples = artifacts$data$data_prepared_tbl %>%
drop_na() %>%
vfold_cv(v = 10)
)
toc()
# 12-month forecast calculations with future dataset
forecast_stacking_tbl <- refit_stacking_tbl %>%
modeltime_forecast(
new_data = artifacts$data$future_tbl,
actual_data = artifacts$data$data_prepared_tbl %>%
drop_na(),
keep_data = TRUE
)
# Toggle OFF parallel processing
plan(sequential)
We invert the Turnover values back to their original values with standardize_inv_vec() and expm1().
Finally, we plot the forecast for the next 12 months within the original scale of the Turnover measure.
lforecasts <- lapply(X = 1:length(Industries), FUN = function(x){
forecast_stacking_tbl %>%
filter(Industry == Industries[x]) %>%
#group_by(Industry) %>%
mutate(across(.value:.conf_hi,
.fns = ~standardize_inv_vec(x = .,
mean = artifacts$standardize$std_mean[x],
sd = artifacts$standardize$std_sd[x]))) %>%
mutate(across(.value:.conf_hi,
.fns = ~expm1(x = .)))
})
forecast_stacking_tbl <- bind_rows(lforecasts)
forecast_stacking_tbl %>%
group_by(Industry) %>%
plot_modeltime_forecast(.title = "Turnover 1-year forecast",
.facet_ncol = 4,
.conf_interval_show = FALSE,
.interactive = TRUE)
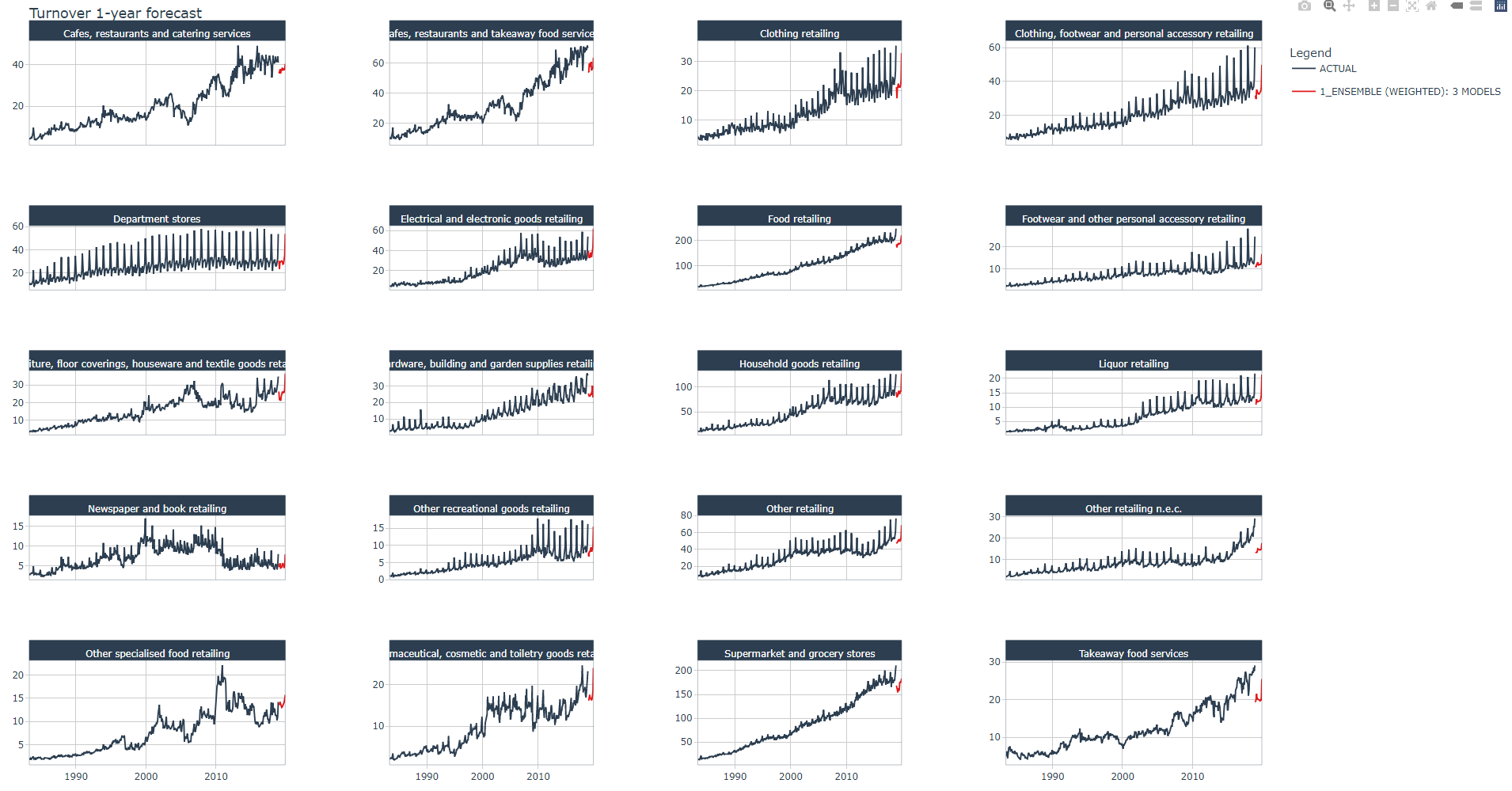
Below a zoomed view of the same plot
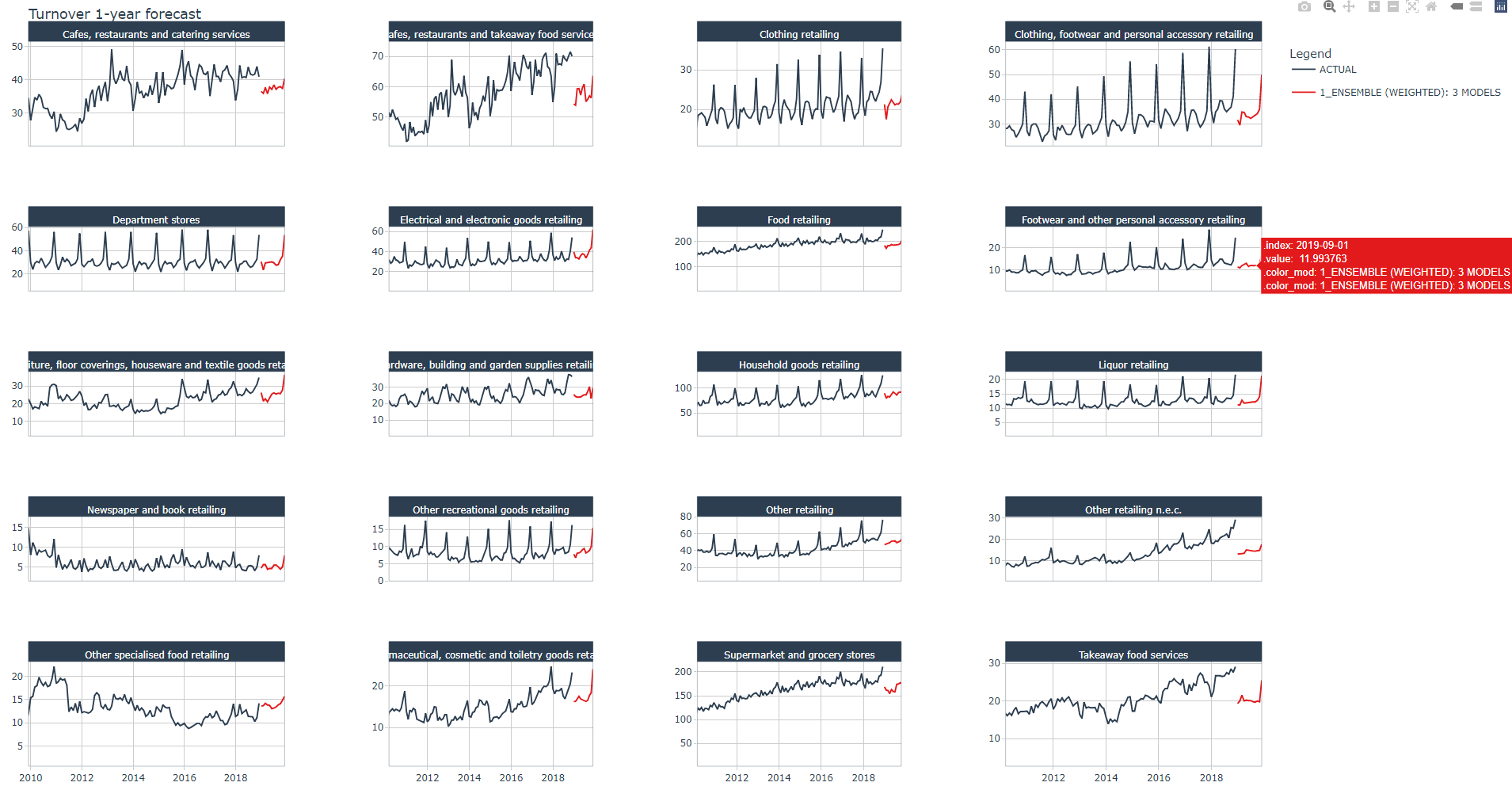
Conclusion
In this article you have learned how to implement stacked ensemble models. A stacking algorithm or meta-learner learns from predictions. The predictions come from k-fold cross-validations applied to each submodel. Once fed into the meta-learner tunable specification, a hyperpameter tuning will be peformed for the meta-learner.
We defined three meta-learners: SVM, Random Forest, and XGBoost from which we defined a weighted average ensemble model (stack-level) using the same method as explained Part 5.
The stacked ensemble algorithm outperformed all algorithms.
References
1 Dancho, Matt, "DS4B 203-R: High-Performance Time Series Forecasting", Business Science University






